Abstract
Deep learning techniques have been incorporated to predict whether a person is affected with lung cancer or not by extracting features from the CT-Scan Images extracted from data repository. these types of predicting the lung cancer system also help us to understand the behavioural patterns present in the images which can helps us to analyse some more information. In this proposed method we give a brief introduction to how we can sue deep learning in medicine and how they might save lives. The deep learning model which was used to train on the images is Convolutional Neural Network, which is a very popular algorithm to work on images.
Algorithm Description
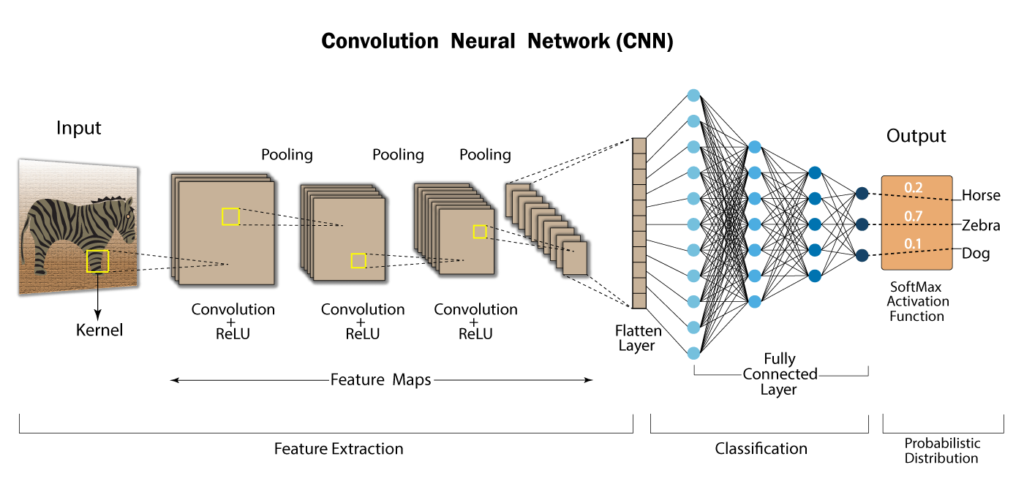
Convolutional Neural Network:
As we all are aware of the fact, how deep learning and transfer learning is revolutionizing the world with its immense capability of handling any kind of data and learning so efficiently. So, similarly we have applied the same concept by picking a deep learning model i.e., Convolutional neural network which basically work son the principle of having filters. Each convolutional layer has some specific filters to identify and extract the features from the input image and learn it and transfer it to other layers for further processing. We can have as many filters as possible in the convolutional layer depending on the data we are dealing on. Filter are nothing but feature detectors in the input data. Along with the convolutional layer we also have other layers which does further pre-processing such as Maxpooling, Activation function, Batch Normalization and dropout layer. These all contribute to the CNN model creation and along with the flatten and output layer. The reason we do flattening is to feed the output of the CNN model to the dense layer which gives us the probability of the predicted value.
How to Execute?
Make sure you have checked the add to path tick boxes while installing python, anaconda.
Refer to this link, if you are just starting and want to know how to install anaconda.
If you already have anaconda and want to check on how to create anaconda environment, refer to this article set up jupyter notebook. You can skip the article if you have knowledge of installing anaconda, setting up environment and installing requirements.txt
- Install the prerequisites/software’s required to execute the code from reading the above blog which is provided in the link above.
- Press windows key and type in anaconda prompt a terminal opens up.
- Before executing the code, we need to create a specific environment which allows us to install the required libraries necessary for our project.
- Type conda create -name “env_name”, e.g.: conda create -name project_1
- Type conda activate “env_name, e.g.: conda activate project_1
- Go to the directory where your requirement.txt file is present.
- cd <>. E.g., If my file is in d drive, then
- d:
7.cd d:\License-Plate-Recognition–main #CHANGE PATH AS PER YOUR PROJECT, THIS IS JUST AN EXAMPLE
8. If your project is in c drive, you can ignore step 5 and go with step 6
9. g., cd C:\Users\Hi\License-Plate-Recognition-main
10. CHANGE PATH AS PER YOUR PROJECT, THIS IS JUST AN EXAMPLE
11. Run pip install -r requirements.txt or conda install requirements.txt (Requirements.txt is a text file consisting of all the necessary libraries required for executing this python file. If it gives any error while installing libraries, you might need to install them individually.)
12. To run .py file make sure you are in the anaconda terminal with the anaconda path being set as your executable file/folder is being saved. Then type python main.pyin the terminal, before running open the main.py and make sure to change the path of the dataset.
13. If you would like to run .ipynb file, Please follow the link to setup and open jupyter notebook, You will be redirected to the local server there you can select which ever .ipynb file you’d like to run and click on it and execute each cell one by one by pressing shift+enter.
Please follow the above links on how to install and set up anaconda environment to execute files.
Data Description
The dataset was downloaded from a kaggle data repository. The dataset has been pre-processed and cleaned to remove any bias while training. Its dataset has been augmented and divided into normal and not normal folders where which each folder consists of around more than 1500 images. Shape of all the images is equally scaled to about 255 x 255 RGB format.
Normal
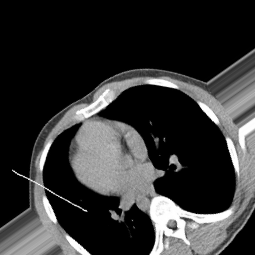
Not Normal
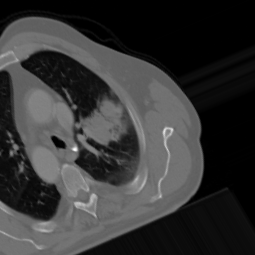
Final Results
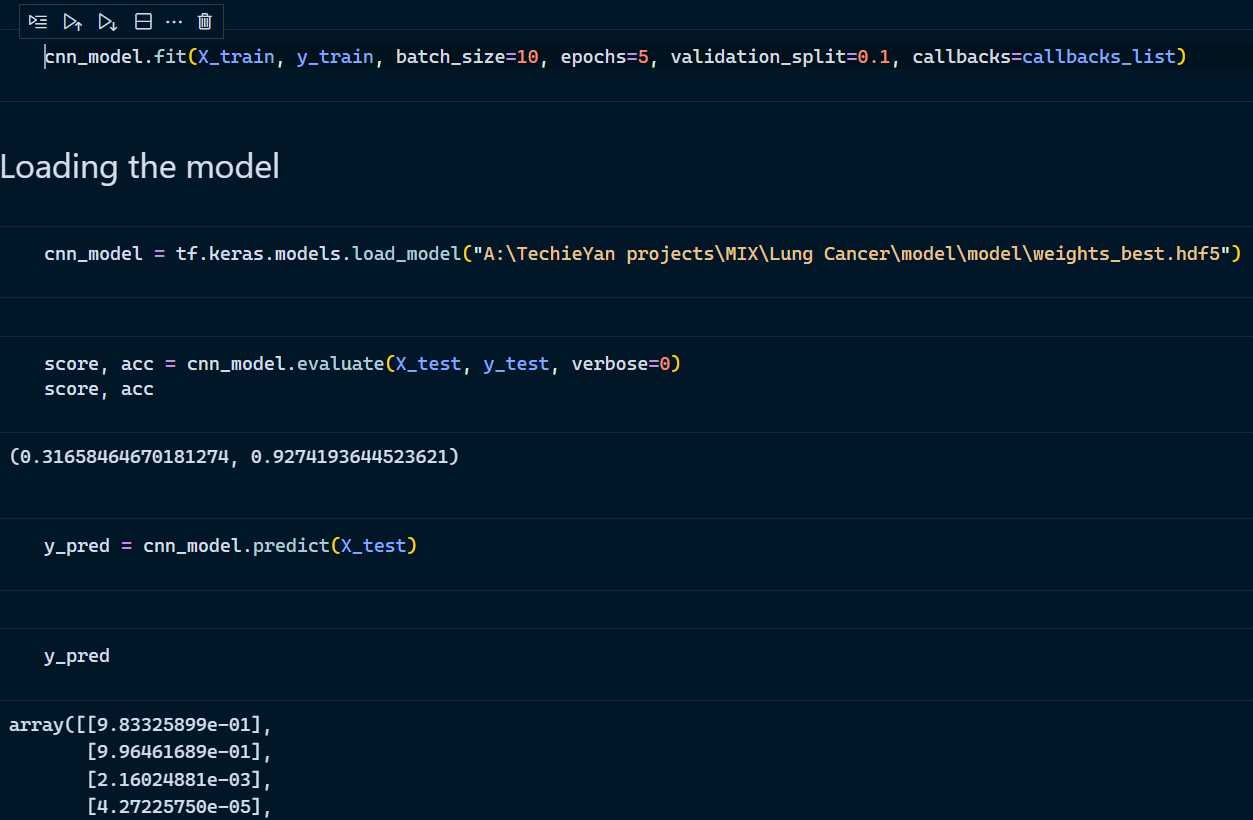
- Model Training and Loading the model
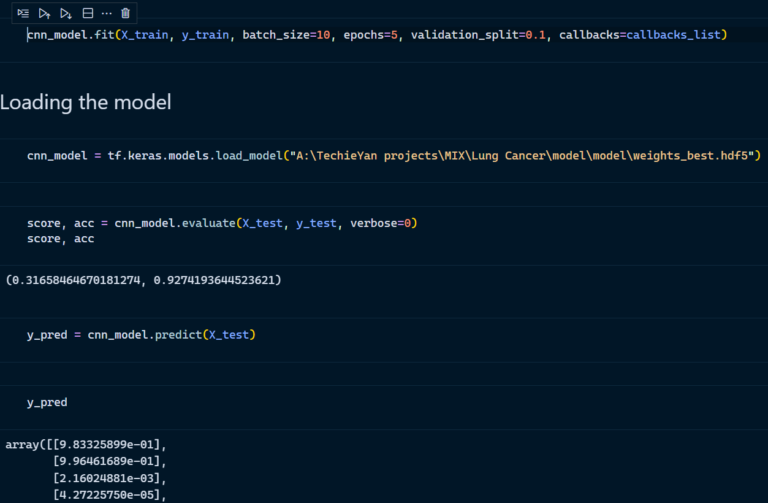
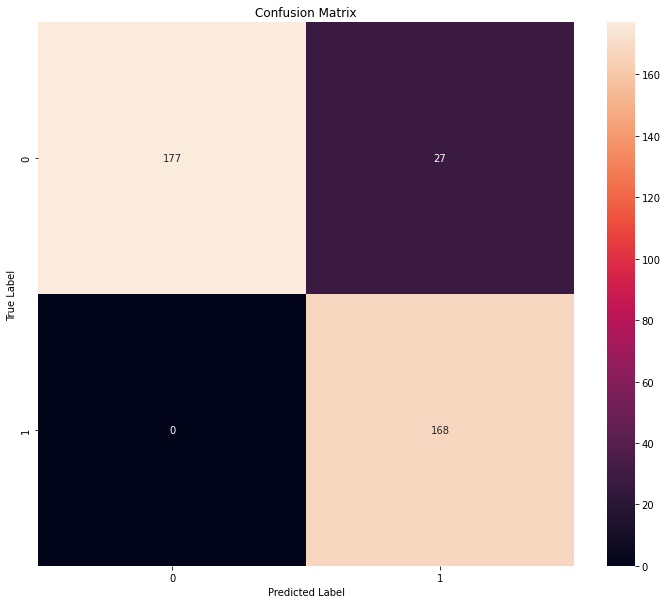
2. Confusion matrix

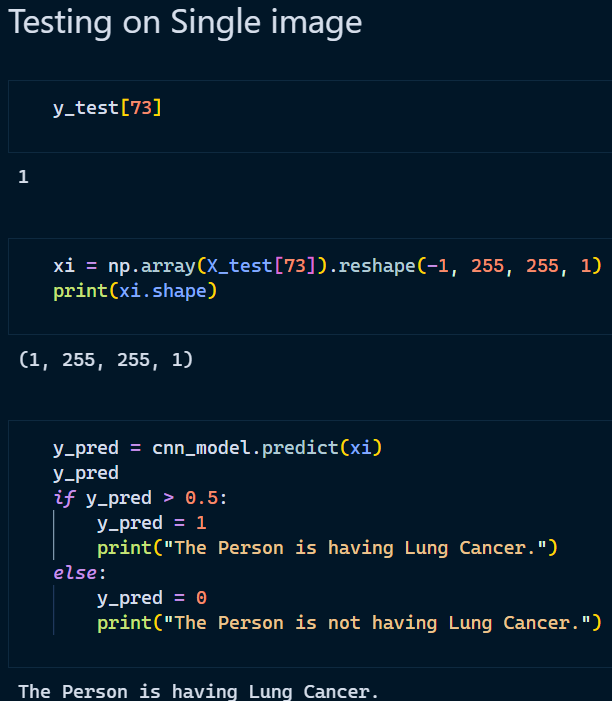
3. Testing on a single image
Evaluation Metrics
Evaluation metrics are considered as one of the most important steps in any machine learning and deep learning projects, where it will allow us to evaluate how good our model is performing on the new data or on unseen data. There are a lot of evaluation metrics which can be used in order to assess how good our predicting the lung cancer model is performing, in our case, since we are dealing with binary classification and neural network, we are going to sue binary_cross_entropy/log_loss, which basically compares the actual class with the predicted probabilities and then it calculates a corrected probability by subtracting it with the probability of a datapoint belonging to class1 with the predicted probability, i.e. for the case of ID8 it is actually class 0, but the probability is of class 1 is 0.56, so we subtract (1 – 0.56), we get 0.44 that is our corrected probability. Then Log_loss is calculated by applying log transformation on each of the calculated_probablities. The the average of the negative corrected_probablities are taken which will gives us the log_loss/binary_cross_entropy, the lower the value the better our predicting the lung cancer model is performing.
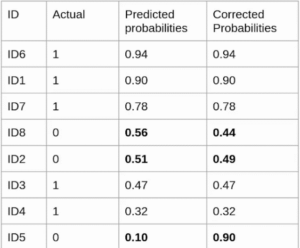
Log_loss calculation for corrected_probablities
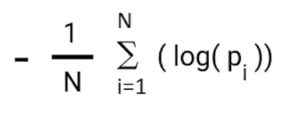
Log_Loss formula without calculating corrected_probablities
![]()
Reference:
Issues you may face while executing the code
- We might face an issue while installing specific libraries, in this case, you might need to install the libraires manually. Example: pip install “module_name/library” i.e., pip install pandas
- Make sure you have the latest or specific version of python, since sometimes it might cause version mismatch.
- Adding path to environment variables in order to run python files and anaconda environment in code editor, specifically in any code editor.
- Make sure to change the paths in the code accordingly where your dataset/model is saved.
Refer to the Below links to get more details on installing python and anaconda and how to configure it.
https://techieyantechnologies.com/2022/07/how-to-install-anaconda/
Note:
All the required data has been provided over here. Please feel free to contact me for model weights and if you face any issues.
Click Here For The Source Code And Associated Files.
https://www.linkedin.com/in/abhinay-lingala-5a3ab7205/
Yes, you now have more knowledge than yesterday, Keep Going.
