Kidney stone detection main Using Transfer Learning
Abstract
Kidney stones are not a new subject but it is one of the major health concerns these days, if not detected at early stages might also become life-threatening. A small piece of stone may pass through urinary track without causing symptoms. If a stone grows to more than 5 millimetres it can cause blockage of the ureter resulting in severe pain in the lower back or abdomen. Hence, it’s necessary to have an approach to detect the stone in the kidney to avoid further health issues. In this project we will be using transfer learning to make the prediction.
Code Description & Execution
Algorithm Description
Resnet50 model:
- This architechture contains (conv1) as first convolutional layer containing in channels as 3 which is due to RGB input tensor , (bn1) as batch normalization layer, followed by ReLU and MaxPooling and then it contains 4 main layers named layer1, layer2, layer3 and layer4 which contains further sub layers of convolution followed by batchnorm followed by relu followed by maxpooling , and then finally fc.
- ReLU activation is used as it’s the most proven activation function for classification problems as it introduces good and right amount of non linearity with less chances of vanishing gradient problem !
- Batch normalization helped in making the network more stable and learning faster thereby faster convergence.
- Maxpooling helped in downsampling high number of parameters created by producing higher dimensional feature maps after convolution operation and thus selecting only relevant features from the high dimensioned feature matrix.
- Then i replaced last layer of this architecture by fully connected layer containing two sub linear layers as follows : Linear(in_features=2048, out_features=512) Linear(in_features=512, out_features=133)
with ReLU activations between the linears.
To read more about ResNet 50 check this link,
https://towardsdatascience.com/understanding-and-coding-a-resnet-in-keras-446d7ff84d33
How to Execute?
So, before execution we have some pre-requisites that we need to download or install i.e., anaconda environment, python and a code editor. Anaconda: Anaconda is like a package of libraries and offers a great deal of information which allows a data engineer to create multiple environments and install required libraries easy and neat.
Download link:
Python: Python is a most popular interpreter programming language, which is used in almost every field. Its syntax is very similar to English language and even children and learning it nowadays, due to its readability and easy syntax and large community of users to help you whenever you face any issues.
Download link:
https://www.python.org/downloads/
Code editor: Code editor is like a notepad for a programming language which allows user to write, run and execute program which we have written. Along with these some code editors also allows us to debug, which usually allows users to execute the code line by line and allows them to see where and how to solve the errors. But I personally feel visual code is very good to work with any programming language and makes a great deal of attachment with user.
Download links:
How to Execute?
Note: Make sure you have added path while installing the software’s.
Install the prerequisites mentioned above.
Step1
Open anaconda prompt and create a new environment. To create an environment use the commands given below. Replace env_name by the name of environment you want to give.
- conda create -n “env_name”
- conda activate “env_name”
Step2
Set up jupyter notebook for your environment
- conda install –c conda-forge jupyterlab
- conda install –c anaconda ipython
Step3
Install necessary libraries from requirements.txt file provided.
Go to the directory where your requirement.txt file is present.
- cd D:\Exicuted Projects\Kidney_stone_detection-main
- Run command pip install -r requirements.txt or conda install requirements.txt
Requirements.txt is a text file consisting of all the necessary libraries required for executing this python file. If it gives any error while installing libraries, you might need to install them individually. All the required files will be downloaded after you run it. I got requirement already satisfied as I already have them installed.
Step4
To run the code, start jupyter notebook by typing “jupytrr notebook” in command prompt, this will navigate directly to jupyter notebook in your default web browser
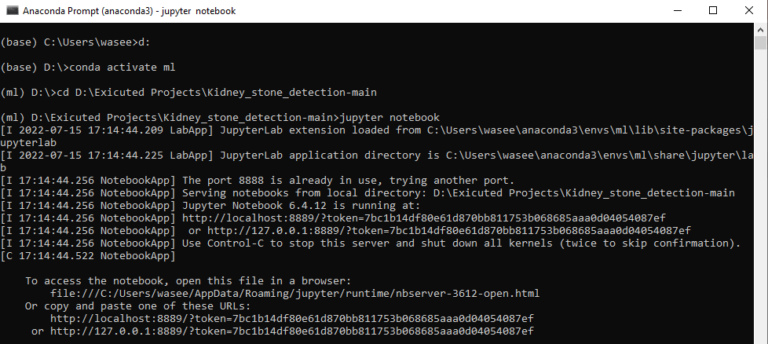
Open the folder containing the code, here it is Kidney stone prediction. When you run the Kidney_stone_detection_Xresnet50.ipynb file, you get the appropriate results.
Data Description
The dataset used is the ct scan images of kidney scans wher total of 1453 images are used of which 1163 images are used for traing and 290 images are used for validating the model and 346 images are used to test the model for different perfomence metrix.
Sample images of dataset
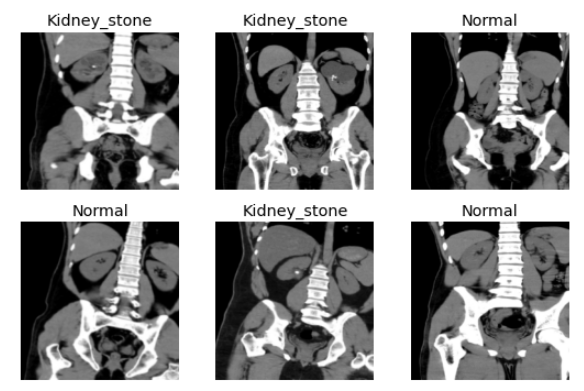
Results
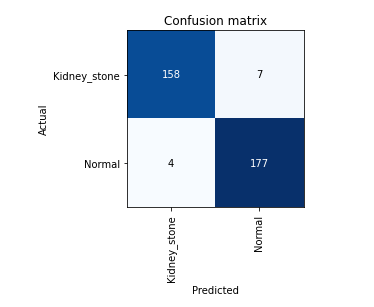
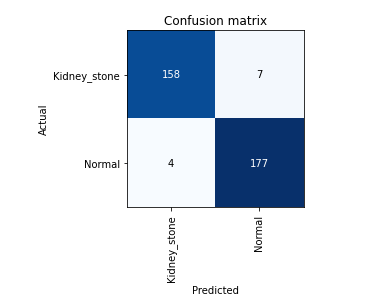
Confusion matrix shows our model accurecy is greater than 96%
Predictions after passing the hidden images to the trained model
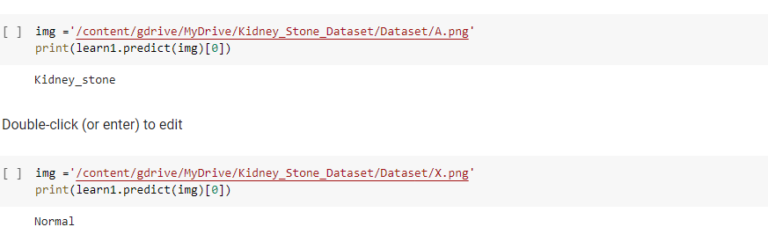
Issues Faced
- Make sure to use .eval()function before making predictions
- Time taken for training is bit longer
Click Here To Download This Code And Associated File.
